10. DNA Packaging (960)
- lscole
- Apr 18, 2025
- 4 min read
Updated: Apr 26
Two human genomes--together totaling about 6 billion base pairs--are packed into the nucleus of every cell in your body. Every cell contains the full genome, even though different cells use different genes depending on their function.
That creates a problem.
If we could stretch out the DNA in a single human cell--laying all 6 billion base pairs end to end--it would extend nearly six feet. Yet the nucleus that holds it is only about 5 micrometers in diameter, roughly 10,000 times smaller than a dot of ink from a ballpoint pen.
Six feet of DNA in a space that small. How is that possible? And more than that--the DNA has to remain usable while it’s packed.
Part of the answer is that DNA is very, very thin—about 2 nanometers across, which is smaller than the wavelength of visible light. But thinness alone isn’t enough. The deeper answer is packaging.
DNA isn't just crammed into the nucleus. It's carefully wound, wrapped and looped. And it's not just about fitting DNA into a small space. DNA packaging plays a major role in determining which genes are available to be transcribed.
Packaging in layers
Packaging doesn’t happen all at once. It builds in layers.
First, DNA wraps around proteins. Then those structures pack together into fibers. Then those fibers loop. And finally, during cell division, everything condenses further.
We’ll look at each of these steps in turn.
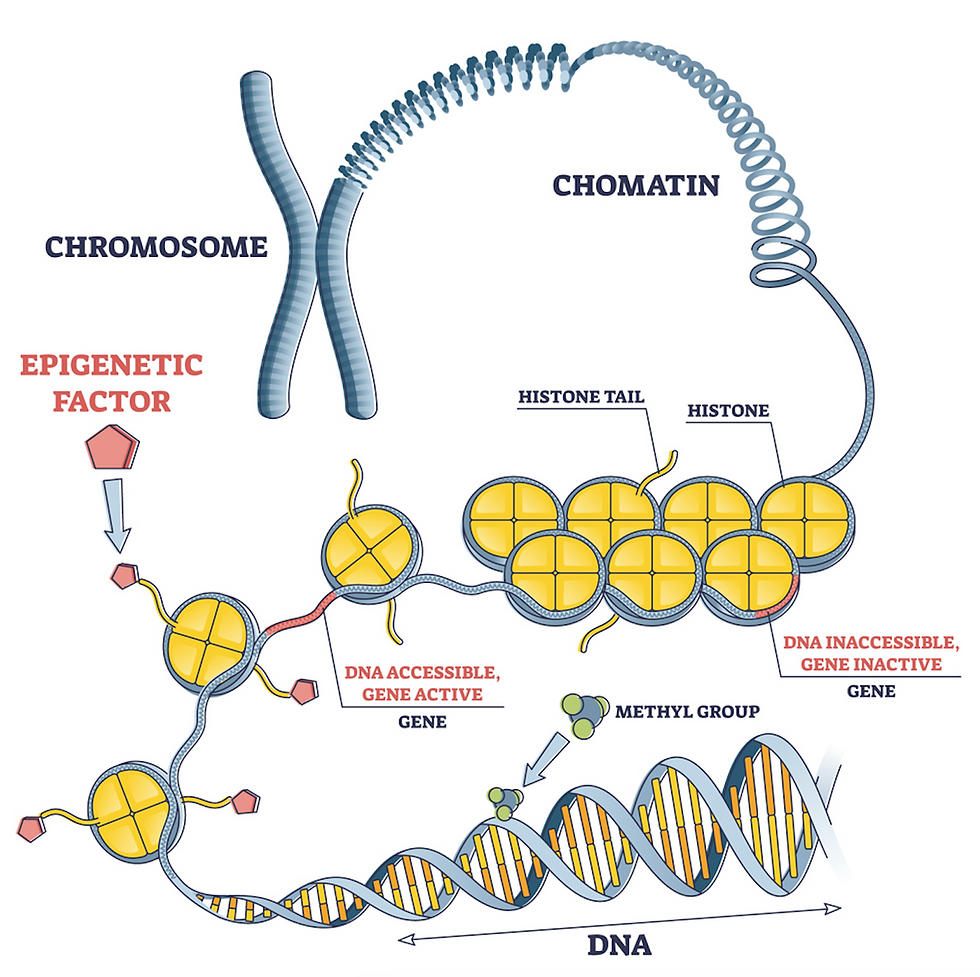
Level 1: Nucleosomes--DNA wrapped around protein spools
In the first level of packaging, DNA wraps around small protein complexes called histones, much like thread wrapped around a spool.
Each nucleosome consists of eight histone proteins, around which about 150 nucleotides of DNA are wound. Between the nucleosomes are short stretches of exposed DNA called linkers.
This combination of DNA and associated proteins is called chromatin. It is often described as looking like “beads on a string.”
It’s tempting to think of this packaging as static. It isn’t.
Nucleosomes are dynamic. They move along the DNA, partially unwrapping and rewrapping. Sections of DNA become briefly exposed--for on the order of milliseconds--allowing other proteins to bind. Specialized protein complexes can also reposition nucleosomes along the DNA.
This constant motion means that even packaged DNA remains accessible.
Level 2: Further compaction
The beads-on-a-string structure is too loose and vulnerable to be safe inside the nucleus. So the beads-on-a-string chromatin is packed more tightly into a fiber.
The exact structure of this fiber is still an area of active research. What’s clear is that interactions between histone proteins--particularly their flexible tails--and a fifth histone (H1) help bring nucleosomes closer together.
The result is a more compact form of chromatin.
Level 3: Looping -- organization and access
At this level, packaging starts to determine which genes the cell can transcribe.
The thick, or compact, chromatin fiber is organized by protein complexes into large loops that contain tens to hundreds of thousands of nucleotides.
These loops are not random.
These loops are not random--they reflect the parts of the genome that need to be accessible. Regions that a cell needs to use--genes that must be expressed--are organized such that they are easier to reach. Other regions are more tucked away.
And, like nucleosomes, these loops are dynamic. They form, dissolve, and reform as the needs of the cell change.
Chemical "signposts" on histones
Another layer of control operates through chemical modifications made to histones.
The histone proteins around which DNA is wrapped have flexible tails that can be chemically modified in different ways. The cell has enzymes that add and remove these modifications.
Sometimes these changes directly affect how tightly DNA is wrapped. But more often, they act as signals--markers that recruit other proteins to that location.
These markers might attract transcription machinery, recruit DNA repair systems, or signal that DNA should be replicated
In this way, histone modifications act like signposts, guiding diverse cellular processes to specific regions of the genome.
Scientists sometimes refer to this system as a histone code--a set of signals that help determine how each stretch of DNA will be used.
Additional layers of control
The cell has other ways of controlling DNA accessibility, too.
In some cases, standard histone proteins are replaced with different versions called histone variants that make the DNA more or less accessible.
In other cases, transcription factors that bind DNA near genes recruit enzymes that modify nearby chromatin, opening or closing access locally.
Even the position of DNA within the nucleus matters. Chromosomes tend to occupy specific regions, known as chromosome territories. Genes that are actively used are generally located toward the interior of the nucleus. Less active regions are more often found closer to the periphery.
A dynamic system
DNA packaging is not static. It's a constantly shifting system. Nucleosomes slide and unwrap. Loops form and then dissolve. Chemical marks are constantly being added and removed. And proteins are also constantly scanning for signals and responding to them.
The genome is not just stored, it is always being reorganized.
Level 4: Maximum compaction during cell division
The final level of packaging occurs during cell division.
As a cell prepares to divide, its DNA becomes dramatically more condensed. The looped chromatin structures are further coiled into tightly packed chromosomes--the familiar X-shaped structures seen in microscope images.
In this form, DNA is no longer accessible for transcription. Gene expression largely pauses or becomes greatly reduced. The priority is not reading the DNA, but safely separating it into two daughter cells.
At every level--from nucleosomes to loops to entire chromosomes--the way DNA is packaged helps determine which genes are available, which are hidden, and when each will be used.
The genome is not just a sequence of letters. It's a dynamic, three-dimensional structure that's constantly being reorganized to meet the needs of the cell.

Comments